My code is reading structured data files and generating Pandoc abstract syntax tree JSON from what it sees.
-
My code is reading structured data files and generating Pandoc abstract syntax tree JSON from what it sees.
I feel like somehow I have upped my text processing pathology by intentionally using Pandoc JSON as an intermediate format
feels good
-
Random Geekreplied to Random Geek last edited by [email protected]
And then I had to let #Nushell drive the #Pandoc batch processing because of course I did.
(tell pandoc to process every JSON file in the cache folder to a corresponding Org file in the out folder)
let $format = "org"
ls cache/*.json | insert target {
$in.name | path parse | upsert extension $format | upsert parent "out" | path join
} | each { pandoc $in.name -o $in.target }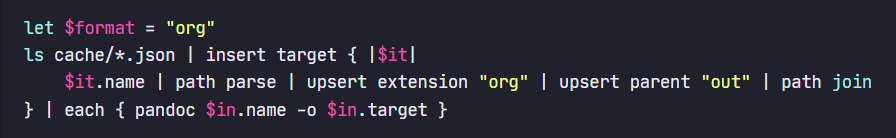
-
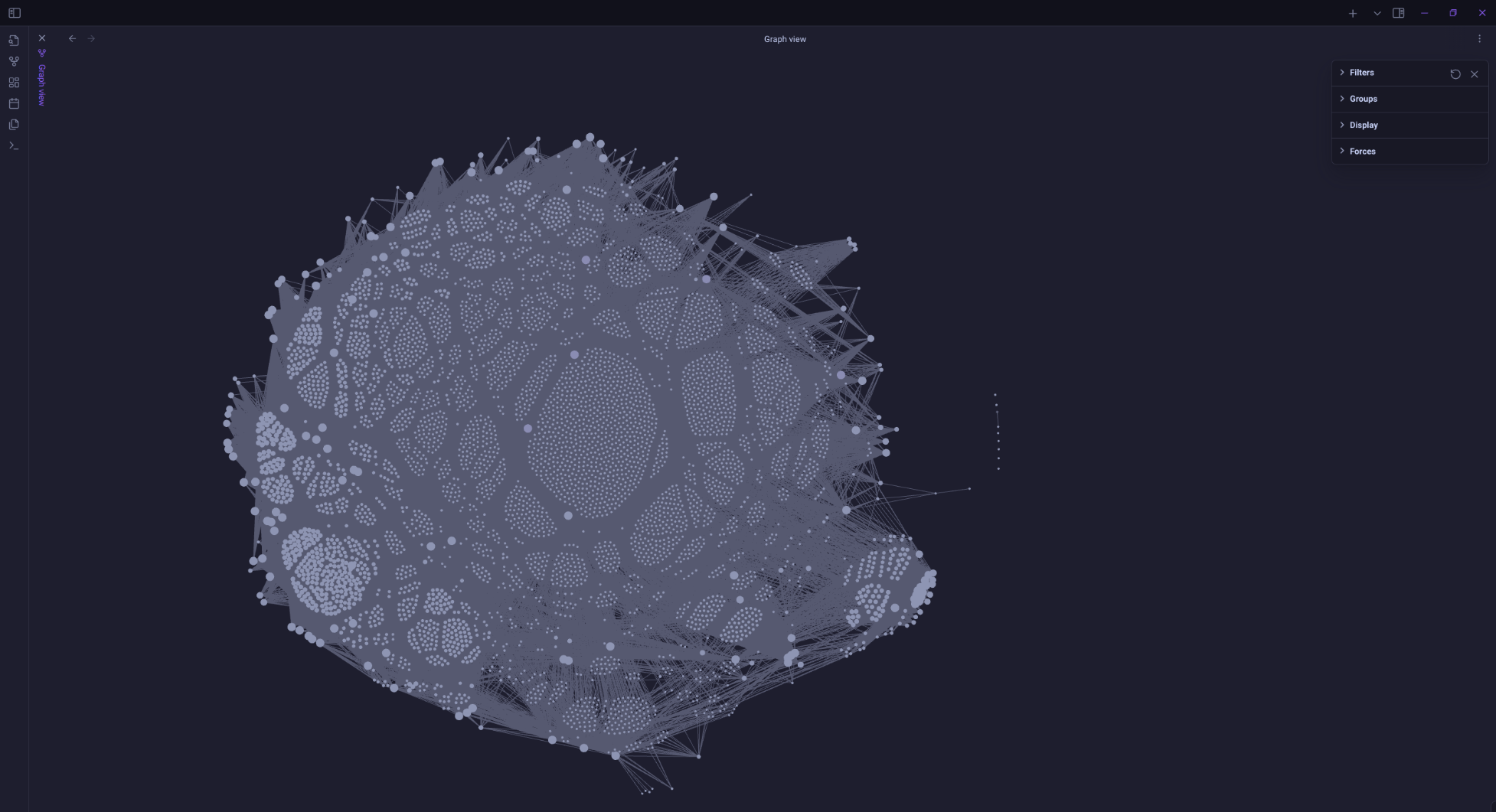
This morning I got basic links between files working.
Probably didn't need to process all 6,100 schema files in that folder.
At least I had the foresight to put output files in a sandbox folder rather than drop them in my main note repo.

-
@randomgeek I‘ve never seen a note graph look like a nanomachine mold colony.
Nice job!
-
@spinningthoughts I don’t think I’ll try replicating this exact process in Logseq. A subset maybe, but even Obsidian took several minutes to index all that.
